Dong Fang’s laboratory published a research article entitled SMYD5 catalyzes histone H3 lysine 36 trimethylation at promoters in Nature Communications on 9th June,2022. It was the first time identified that SMYD5, except SETD2, could act as a methyltransferase of H3K36me3 in mammalian cells, catalyzing and accumulating H3K36me3 at promoters through screening (figure 1).
As carriers of epigenetic information, histone modifications play important roles in regulating gene expression. H3K36me3, which has been widely studied, is generally considered the active marker enriched in intergenic regions and gene bodies. H3K36me3 participates in cellular activities, DNA damage, regulation of transcription initiation, RNA splicing, m6A RNA modification et al.
In previous studies, the traditional method ChIP-seq found that H3K36me3 is mainly enriched in the active gene bodies and gradually increased from the 5’ to 3' end. H3K36 methyltransferase was originally identified as Set2 in yeast1, and its homolog protein SETD2 was identified as H3K36me3 methyltransferase in mammalian cells. In addition, the meiotic-specific histone methyltransferase PRDM9 has been reported to catalyze both H3K4me3 and H3K36me3 in testis2.
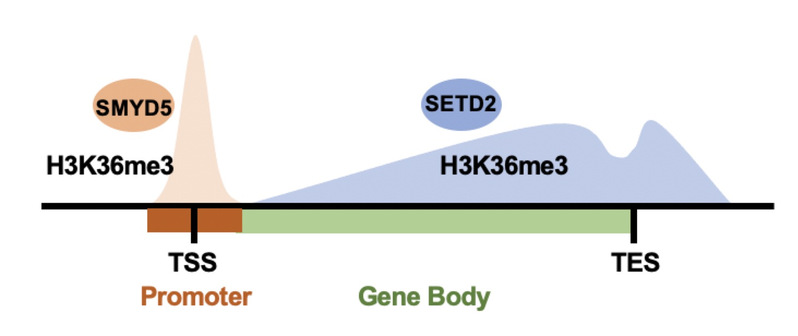
Figure 1. SMYD5 methylates H3K36me3 at promoters, while SETD2 methylates H3K36me3 at gene body regions.
In this study, the authors use the method CUT&Tag to analyze the distribution of H3K36me3 in uncross-linked cells, by which Tn5 transposons were used to cut and label DNA as natural as possible. Surprisingly, H3K36me3 was found deposited onto chromatin at the gene promoters for the first time. This enrichment has also been validated in native H3K36me3 ChIP-seq without sonication. Through a targeted gene screening, SMYD5 was identified as a methyltransferase to catalyze the methylation of H3K36me3 in vivo and in vitro.
SMYD5 is recruited to chromatin by RNA polymerase II, resulting in the enrichment of H3K36me3 at promoters. The C-terminal glutamate-rich domain of SMYD5 is important for its binding with histone H3 and methylation of H3K36me3.
In addition, elevated Smyd5 expression contributes to the liver hepatocellular carcinoma (LIHC) tumorigenesis. Knockdown SMYD5 can effectively inhibit the occurrence of LIHC in the mouse model. Overexpression of wild-type Smyd5 in knockdown mice can rescue the tumorigenesis, while overexpression of Smyd5 with deleted C-terminal glutamate-rich domain could not.
Taken together, these data provide a new idea for further exploring the function of H3K36me3 and understanding the abnormal H3K36me3 in tumorigenesis.
Dong Fang is the corresponding author of this article. Yanjun Zhang, Yuan Fang and Yin Tang, from Life Sciences Institute, Zhejiang University, are the co-first authors. This work is supported by Professor Bin Zhao on the mouse LIHC model. This work is also supported by the National Natural Science Foundation of China.
References
1. Strahl, B.D. et al. Set2 is a nucleosomal histone H3-selective methyltransferase that mediates transcriptional repression. Mol Cell Biol 22, 1298-1306 (2002).
2. Powers, N.R. et al. The Meiotic Recombination Activator PRDM9 Trimethylates Both H3K36 and H3K4 at Recombination Hotspots In Vivo. PLoS Genet 12, e1006146 (2016).